news

news
Bob Gotwals, NCSSM Instructor of Chemistry and pioneer of the school’s computational science program, wrote this summary about methods and tools of scientific visualization that are used by students in NCSSM’s computational science courses, for the NCSSM Online Connection, the school’s newsletter for NCSSM-Online students and their families:
“A picture is worth a thousand words,” so the saying goes. Students in the NCSSM Computational Sciences program – the majority of which (nine of 11 courses) are offered through the NCSSM-Online program – are embracing Frederick R. Barnard’s advertising advice (taken from Chinese philosophers), published in the December 1921 edition of Printers’ Ink.
Everyone is familiar with standard ways of representing data – scatterplots with the well-utilized X-Y axes, the occasional bar chart or histogram, the oftentimes misleading pie charts. Basic graphing techniques and tools are used in a variety of classes, primarily science and math, but sometimes in other venues, including humanities.
Computational science students, on the other hand, are using the technologies, techniques, and tools of scientific visualization – a way of rendering large amounts of scientific data into sophisticated 2D and 3D graphical representations. In scientific visualization, data is collected from a variety of sources, including lab instruments, remote sensors, and, in our case, from computation model runs. Once data is obtained, a variety of filtering techniques are typically applied, allowing the scientists to use only that data that is relevant to the problem of interest.
While students learn how to use tools such as Microsoft Excel spreadsheets to do simple visualizations, in most of the courses, more advanced software is used.
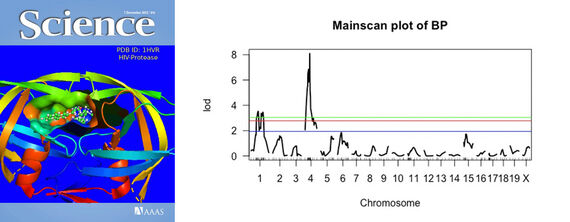
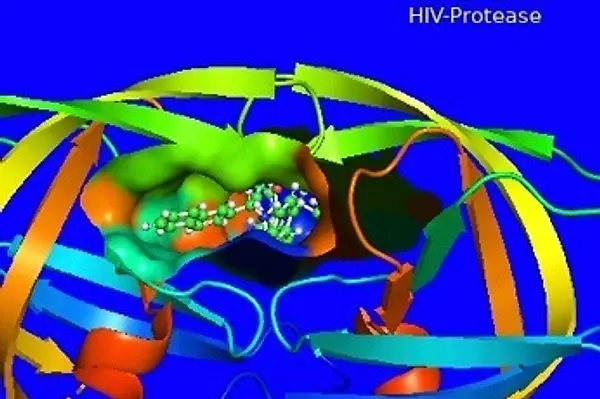
For example, in “Computational Biology/Bioinformatics,” students learn how to visualize 3D representations of proteins, with filters to focus on a specific part of the protein (such as the chains), or to create surface plots of where a drug might bind to that protein. During the Spring 2021 Online Weekend, bioinformatics students selected a protein, generated a graphic, and prepared their visualization as it might appear on the front cover of Science magazine [GRAPHIC: ScienceCover2]. Bioinformatics students also used specialized graphics tools to analyze DNA data from mice. The graphic “Mainscan Plot of BP” shows the mice chromosomes on the x-axis and then a plot of genetic effects on blood pressure. Note that Chromosomes 1 and 4 are both significant, suggesting that the genes controlling blood pressure can be found there [GRAPHIC: MainScan].
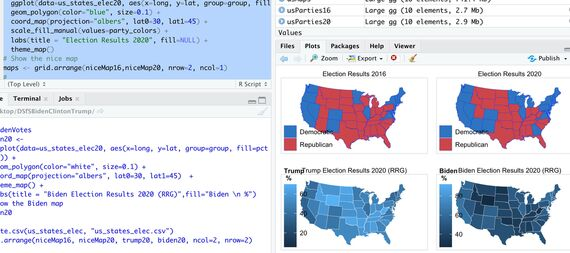
Students in “Data Science for Science” learned how to use specialized graphics packages such as tidyverse (the software name is lowercase) to analyze interesting datasets. One of them was creating maps of electoral college votes for the 2016 and 2020 presidential elections [GRAPHIC: ElectionResults]. Learning how to visualize data represented by various types of mapping tools is a critical skill in the field of data science. Students in the “Research Experience in Computational Science” course used an advanced tool from the Department of Energy, Paraview, to render 3D MRI images of a patient under anesthesia (note the endotracheal tube!) [GRAPHIC: HeadshotParaview]
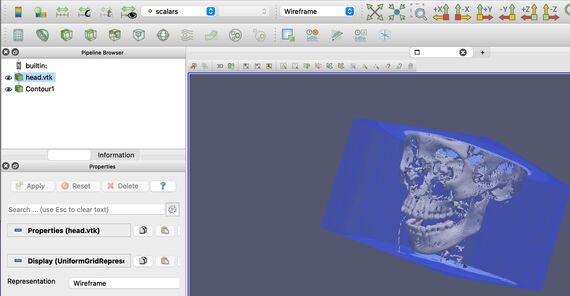
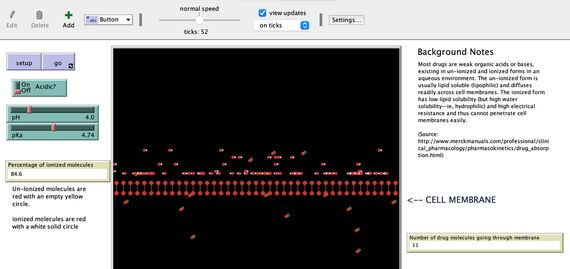
Finally, at the most recent online weekend, participants learned how to build agent-based models using NetLogo. One of the models built, specifically targeting the students in “Medicinal Chemistry,” shows different types of drug molecules going through a phospholipid membrane. This interactive, visualization-based computer model allows those students to explore different contributing factors to the transport of drugs in the body [GRAPHIC: NetLogoDrugMigration].
The scientific visualization skills that computational students are developing will be in great demand in a university research lab, and we’re excited to have access to the research-grade tools that allow students to develop these skills.
You can read more about the computational science program here.